J.S. Garavelli, Hongzhan Huang and D.J. Miller
Protein Information Resource
National Biomedical Research Foundation
Washington, DC 20007
Poster presented at the Protein Society Meeting, August 5 - 9, 2000, San Diego, CA
Abstract
The reported structure of the "putative prismane" protein from Desulfovibrio vulgaris
indicates that it contains a hybrid [4Fe-2S-3O] cluster ligated by seven protein residues,
three cysteines, one persulfido-cysteine, two glutamates and one histidine. The sequences
of at least six other closely homologous proteins have been found, five of these from complete
genomes. In three sequences, the hybrid cluster is associated with an amino terminal [4Fe-4S]
cluster, while in the sequence from Escherichia coli it is associated with a [2Fe-2S]
cluster. In the sequence from Methanobacterium thermoautotrophicum, it may be associated
with a rubredoxin-type center. The nickel-containing chain of carbon-monoxide dehydrogenase
also has a domain with some sequence similarities to the hybrid cluster proteins. The
nickel-containing cluster in carbon-monoxide dehydrogenase may be structurally related to the
hybrid [4Fe-2S-3O] cluster. Models have been prepared for the hybrid [4Fe-2S-3O] and the nickel
hybrid clusters in the RESID Database, distributed by the Protein Information Resource (PIR)
and made available on the Website at
| http://pir.georgetown.edu/pirwww/dbinfo/resid.html. |
The RESID Database is supported by NSF grant DBI-9808414.
Introduction
An unusual iron-sulfur containing protein from Desulfovibrio vulgaris (Hildenborough) was reported in 1992. Its unusual redox behavior and electron paramagnetic resonance spectra led to the prediction that the protein contained a symmetric 6Fe-6S cluster, and it was consequently called the "prismane protein" [1,2]. However, in 1998 a 1.7 Ångstrom resolution X-ray structure for the protein demonstrated that it did not have a 6Fe-6S cluster but two different 4Fe clusters [3]. One was a 4Fe-4S cubane cluster bound by a novel Cys-2X-Cys-8X-Cys-5X-Cys motif. The other cluster was a relatively open 4Fe cluster with a mix of sulfur and oxygen bridging atoms bound by seven amino acid residues including three cysteines, two glutamates, one histidine and a stable persulfido-cysteine, previously seen only as an intermediate active site species (RESID:AA0269). Although the protein was isolated while attempting to purify a periplastic hydrogenase complex, its activity and metabolic role remain unclear. Homologs of the hybrid cluster protein are found in a number of bacterial and archaebacterial species and a division into three classes has been proposed [4].
In 1999 we observed that the hybrid cluster protein shared a distant sequence similarity with the beta chain of carbon-monoxide dehydrogenase found in some bacterial and archaebacterial species. This similarity was also observed by W. A. M van den Berg and others and reported earlier this year [4]. The carbon-monoxide dehydrogenase beta chain is known to contain both nickel and iron but has no observable sequence similarity with other nickel-containing proteins [5,6]. Using the reported structure of the hybrid cluster protein and its sequence similarity with the carbon-monoxide dehydrogenase beta chain, we sought to construct a model for the metallic cofactor as a hybrid nickel-iron cluster, and determine whether there were genes for the proteins that would be required to synthesize this complex cofactor.
Alignment of Homologs
Using ClustalW [7] to align 7 hybrid cluster proteins and 2 of 4 carbon-monoxide dehydrogenase beta chains, we found that they most significantly share sequence similarity in a 280 to 350 residue region at their carboxyl ends. This region includes two folding domains and all seven of the cofactor binding residues observed in the Desulfovibrio vulgaris structure. Those seven residues are conserved in all the hybrid cluster proteins. This region is an homology domain that is observed in at least four homeomorphic superfamilies [8] and is presented as an alignment in Figure 1. A hidden Markov model constructed using this alignment found no other sequences with significant similarity beyond those already observed [9]. The carbon-monoxide dehydrogenase beta chain sequences contain short insertions relative to the hybrid cluster proteins. They also have conservative variations at two cofactor binding positions. Compared with the Desulfovibrio vulgaris hybrid cluster protein, the sequences from Clostridium thermaceticum and Rhodospirillum rubrum have serine rather than glutamate at the seventh binding site (residue 494).
All the proteins have at least one region with four close but variably spaced cysteine residues in front of the hybrid cluster homology domain. Four hybrid cluster proteins have a poorly conserved region of about 120 residues inserted between the amino terminal four-cysteine region and the hybrid cluster homology domain. The hybrid cluster protein from Methanobacterium thermoautotrophicum has an additional region before the four-cysteine region. As originally translated from the whole genome this sequence had the carboxyl half of a strong rubredoxin homology at its amino terminal end [10], an anomaly not remarked upon by van den Berg and others in their classification [4]. When the translation was extended, a more appropriate initiation site was found 15 codons before the reported initiation, and the revised sequence has a complete rubredoxin homology domain. This revised sequence is available in the PIR-International Protein Sequence Database as entry PIR:A59199 [11].
Hybrid Cluster Models
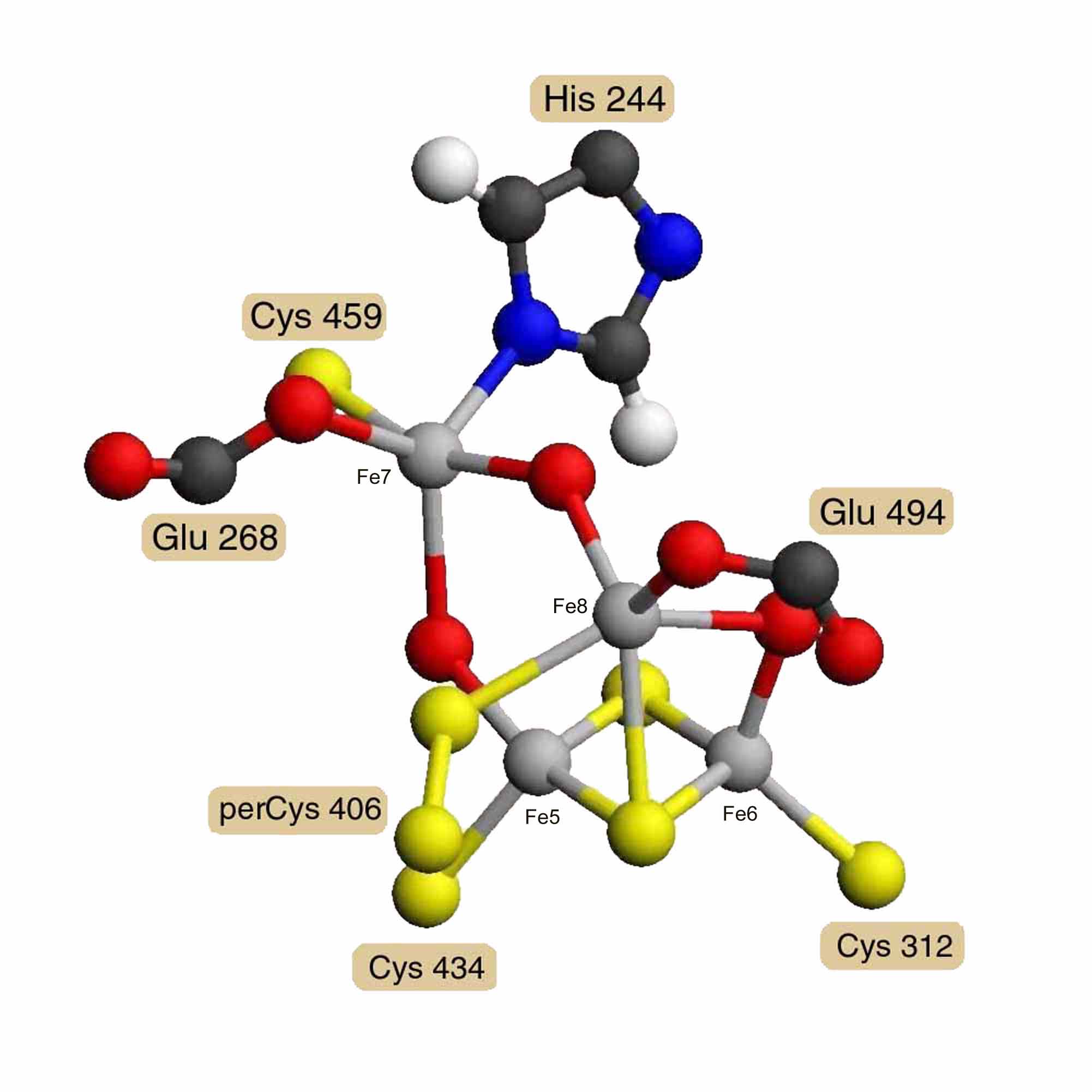
A model of Desulfovibrio vulgaris hybrid cluster [4Fe-2S-3O] was built using the diagrams and distances reported by A. F. Arendsen and others [3]. Model distances and angles were optimized and steric hindrance was reduced using the Alchemy 2000 program [12]. Energy minimization was not attempted for the unparameterized metal clusters. Figure 2 presents a diagram of this model. Coordinates in PDB format are available as entry AA0268 in the RESID Database. The hybrid cluster is built on a 2Fe-2S cluster bound by cysteines 312 and 434. A third iron Fe8 has 5 ligands: a m3 bridge sulfur of the 2Fe-2S cluster, a bridge oxygen to iron Fe6 of the 2Fe-2S cluster, glutamate 494, cysteine persulfide 406, and a bridge oxygen to the fourth iron Fe7. Iron Fe7 is also bound by glutamate 268, cysteine 459, and the N1' of histidine 244. The atom shown as a m2 bridge oxygen connecting Fe7 and Fe5 of the 2Fe-2S cluster is an unidentified atom with partial occupancy and probably constitutes the active site.
Assuming that the carbon-monoxide dehydrogenase beta chain is homologous, a model for its nickel/iron/sulfur cofactor was built as a hybrid [Ni-3Fe-2S-3O] cluster based on the hybrid [4Fe-2S-3O] cluster model. X-ray absorption spectrographic studies of the carbon-monoxide dehydrogenase beta chain of Rhodospirillum rubrum [13] indicate that the nickel is probably ligated by 2 sulfur atoms and 2 or 3 nitrogen or oxygen atoms. This means that the iron atoms labeled Fe5 and Fe6 would probably not be replaced by nickel, since they each have 3 sulfur ligands. The iron labeled Fe7 has 1 sulfur, 1 nitrogen, 2 oxygen and one other undetermined ligand, possibly carbon monoxide. The iron labeled Fe8 has 2 sulfur and 3 oxygen atom ligands, one of which would be from a serine residue rather than a glutamate in the Rhodospirillum rubrum sequence. While Fe7 cannot be ruled out if the undetermined bridge atom were a sulfur, Fe8 meets all the experimental criteria for replacement by nickel, and the model was constructed on this assumption. Coordinates for two alternatively ligated forms of the hybrid [Ni-3Fe-2S-3O] cluster model are available as entries RESID:AA0292 and RESID:AA0293.
Discussion
Sequence alignment of the of the hybrid cluster [4Fe-2S-3O] proteins and carbon-monoxide dehydrogenase beta chains shows that five of the seven cluster ligands, including a novel cysteine persulfide residue, are conserved in both sets of proteins. The seventh ligand is a glutamate in the hybrid cluster proteins and either a glutamate or a serine in the carbon-monoxide dehydrogenase beta chains. The sequence context of the seventh ligand residue is conserved in the mutual alignment, and both the experimental data and the cofactor model accept this replacement. The second ligand is a glutamate residue that is conserved in each of the two sets of proteins but not in the context of the mutual alignment. Plausible alignments can be produced placing other glutamic acid residues, aspartic acid or cysteine residues in this ligand position. However, overall the conservation of the ligand residues supports the hypothesis that hybrid cluster [4Fe-2S-3O] proteins and carbon-monoxide dehydrogenase beta chains are probably homologous and have structurally similar hybrid metal clusters.
The iron-sulfur clusters are produced through the action of the homologs of nitrogenase nifS and nifU proteins. The cysteine desulfurase nifS removes sulfur from free cysteine, modifying an internal cysteine residue to a cysteine persulfide intermediate before contributing the sulfur to a transient [2Fe-2S] iron-sulfur cluster in the iron-sulfur cofactor synthesis protein nifU. No close homologs of these genes were found among the genes likely to be expressed within the same operons as the hybrid cluster protein and carbon-monoxide dehydrogenase beta chain genes. Homologs of nifS have been found in all six organisms with completely sequenced genomes and having either hybrid cluster proteins or carbon-monoxide dehydrogenase beta chain. However, no close homologs of nifU have yet been identified in the archaebacteria Methanococcus jannaschii, Methanobacterium thermoautotrophicum or Pyrococcus abyssi. It might also be expected that genes for proteins that would support the selective incorporation of nickel should be found in the organisms with the hybrid [Ni-3Fe-2S-3O] cluster.
Acknowledgments
We thank Joseph Janda and Kali C. Lewis for administrative support. The RESID database of protein
structure post-translational modifications is supported by NSF grant DBI-9808414. The RESID Database,
accessible on the web at
| http://pir.georgetown.edu/pirwww/dbinfo/resid.html, |
This presentation is available at
| http://pir.georgetown.edu/pirwww/aboutpir/doc/ps00jsgfl.html. |
Table 1. RESID Database
Click here for current database list.
Figure 2. Hybrid [4Fe-2S-3O] Cluster

Model of the Desulfovibrio vulgaris hybrid cluster [4Fe-2S-3O] based on the diagrams and distances of A. F. Arendsen, and others, derived from 1.7 Å resolution X-ray crystallography [3]. What is shown as a m2 oxygen connecting Fe7 and Fe5 is an unidentified atom with partial occupancy and may be the active site.
Figure 3. Proposed Hybrid [Ni-3Fe-2S-3O] Cluster
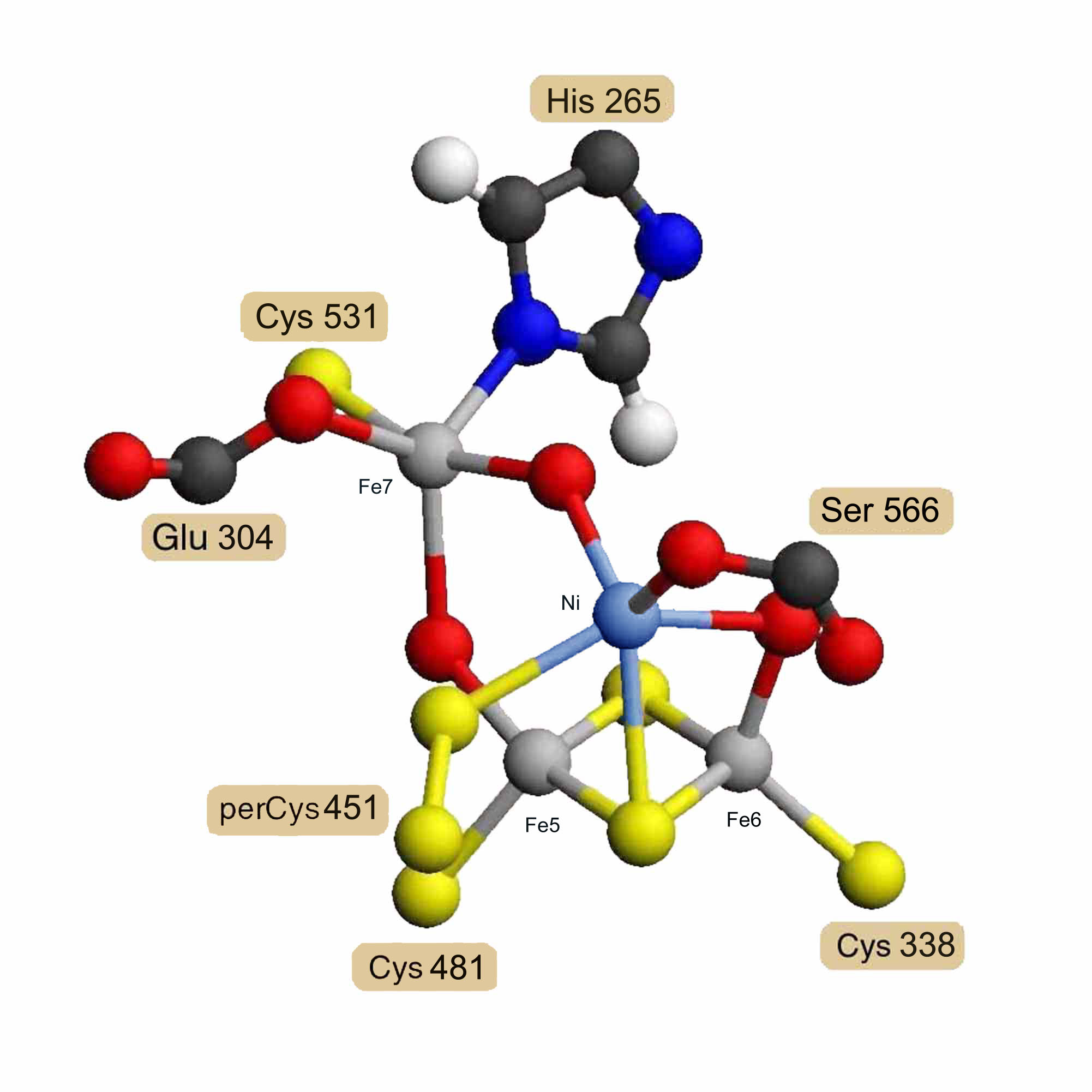
Model for the Rhodospirillum rubrum carbon-monoxide dehydrogenase beta chain nickel-containing cluster as a hybrid [Ni-3Fe-2S-3O] cluster based on homology and the structure of the hybrid [4Fe-2S-3O] cluster.
References
[1] Pierik, A. J., Wolbert, R. B. G., Mutsaers, P. H. A., Hagen, W. R., and Veeger, C. (1992). Purification and biochemical characterization of a putative [6Fe-6S] prismane-cluster-containing protein from Desulfovibrio vulgaris (Hildenborough). Eur. J. Biochem. 206, 697-704.
[2] Pierik, A.J., Hagen, W.R., Dunham, W.R., and Sands, R.H. (1992). Multi-frequency EPR and high-resolution Mössbauer spectroscopy of a putative [6Fe-6S] prismane-cluster-containing protein from Desulfovibrio vulgaris (Hildenborough). Eur. J. Biochem. 206, 705-719.
[3] Arendsen, A. F., Hadden, J., Card, G., McAlpine, A .S., Bailey, S., Zaitsev, V., Duke, E. H. M., Lindley, P. F., Kröckel, M., Trautwein, A. X., Feiters, M. C., Charnock, J. M., Garner, C. D., Marritt, S. J., Thomson, A. J., Kooter, I. M., Johnson, M. K., van den Berg, W. A. M., van Dongen, W. M. A. M., and Hagen, W. R. (1998). The "prismane" protein resolved: X-ray structure at 1.7 Å and multiple spectroscopy of two novel 4Fe clusters. J. Biol. Inorg. Chem. 3, 81-95.
[4] van den Berg, W. A. M., Hagen, W. R., and van Dongen, W. M. A. M (2000). The hybrid-cluster protein ('prismane protein') from Escherichia coli: Characterization of the hybrid cluster protein, redox properties of the [2Fe-2S] and [4Fe-2S-2O] clusters and identification of an associated NADH oxidoreductase containing FAD and [2Fe-2S]. Eur. J. Biochem. 267, 666-676.
[5] Kerby, R. L., Hong, S. S., Ensign, S. A., Coppoc, L. J., Ludden, P. W.; and Roberts, G. P. (1992). Genetic and physiological characterization of the Rhodospirillum rubrum carbon monoxide dehydrogenase system. J. Bacteriol. 174, 5284-5294.
[6] Tan, G. O., Ensign, S. A., Ciurli, S., Scott, M. J., Hedman, B., Holm, R. H., Ludden, P. W., Korszun, Z. R., Stephens, P. J., and Hodgson, K. O. (1992). On the structure of the nickel/iron/sulfur center of the carbon monoxide dehydrogenase from Rhodospirillum rubrum: an X-ray absorption spectroscopy study. Proc. Natl. Acad. Sci. USA 89, 4427-4431.
[7] Thompson, J. D., Higgins, D. G., and Gibson, T. J. (1994). CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673-4680.
[8] George, D.G. (1993) Proposal for the Definition of a "Protein Superfamily". National Biomedical Research Foundation, Washington, DC, pp.1-13. http://pir.georgetown.edu/pirwww/otherinfo/sfdef.html
[9] Krogh, A., Brown, M., Mian, I. S., Sjolander, K., and Haussler, D. (1994). Hidden Markov models in computational biology: Applications to protein modeling. J. Mol. Biol. 235, 1501-1531.
[10] Smith, D. R., Doucette-Stamm, L. A., Deloughery, C., Lee, H., Dubois, J., Aldredge, T., Bashirzadeh, R., Blakely, D., Cook, R., Gilbert, K., Harrison, D., Hoang, L., Keagle, P., Lumm, W., Pothier, B., Qiu, D., Spadafora, R., Vicaire, R., Wang, Y., Wierzbowski, J., Gibson, R., Jiwani, N., Caruso, A., Bush, D., Safer, H., Patwell, D., Prabhakar, S., McDougall, S., Shimer, G., Goyal, A., Pietrokovski, S., Church, G. M., Daniels, C. J., Mao, J., Rice, P., Noelling, J., and Reeve, J. N. (1997). Complete genome sequence of Methanobacterium thermoautotrophicum Delta H: functional analysis and comparative genomics. J. Bacteriol. 179, 7135-7155.
[11] Barker, W. C., Garavelli, J. S., Huang, H., McGarvey, P. B., Orcutt, B. C., Srinivasarao, G. Y., Xiao, C., Yeh, L-S. L., Ledley, R. S., Janda, J. F., Pfeiffer, F., Mewes, H-W., Tsugita, A., and Wu, C. (2000). The Protein Information Resource (PIR). Nucleic Acids Res. 28, 41-44.
[12] Tripos, Inc. St. Louis, MO 63144. http://www.tripos.com
[13] Tan, G. O., Ensign, S. A., Ciurli, S., Scott, M. J., Hedman, B., Holm, R. H., Ludden, P. W., Korszun, Z. R., Stephens, P. J., Hodgson, K. O. (1992). On the structure of the nickel/iron/sulfur center of the carbon monoxide dehydrogenase from Rhodospirillum rubrum: an x-ray absorption spectroscopy study. Proc. Natl. Acad. Sci. USA 89, 4427-4431.
[14] Garavelli, J.S. (1999) The RESID database of protein structure modifications. Nucleic Acids Res. 27, 198-199.